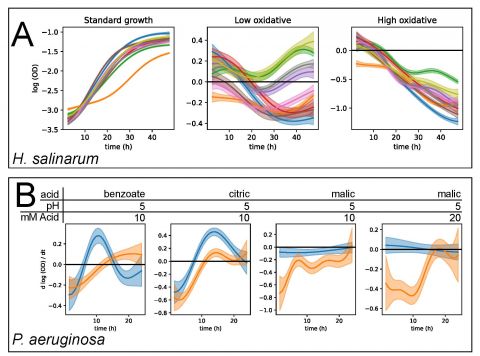
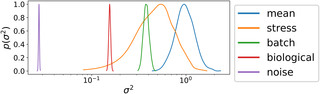
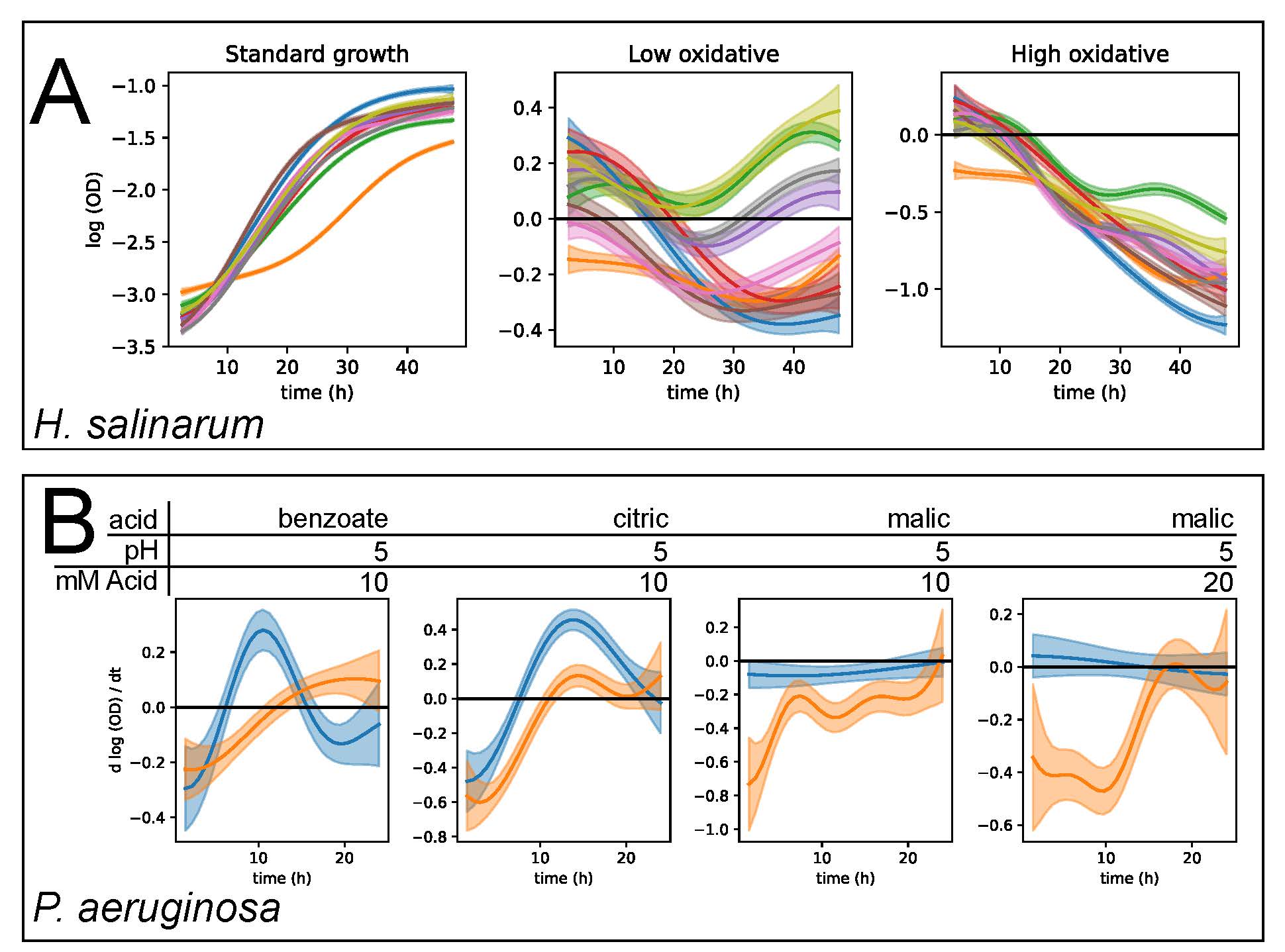
The goal of this Data+ project is to apply and extend custom analytics solutions to understand and predict microbial population growth.An explosion of data has resulted from tracking the growth of bacteria in high throughput devices. These data were generated to understand how microbes grow. Better models that fit and predict these growth data are needed for better treatment of pathogenic bacterial infections, food safety, beer and bread fermentation, and understanding stress resilience of the microbiome. Using nonparametric statistical models to analyze how microbes grow under stress, the Schmid research lab at Duke has made important discoveries in these areas. These studies generated large data sets and developed statistical models to track and predict how microbes grow and change their gene expression when faced with extreme stress. We built a web application called phenom to make these models accessible to the broader community. In this Data+ project, students will beta test the web app and make improvements, including data visualization, extending the underlying statistical model, and analyzing data using the app.
Project Lead: Amy Schmid
Project Manager: Andrew Soborowski
Image credit: Tonner, P.D., Darnell, C.L., Bushell, F.M.L., Lund, P.A., Schmid, A.K.*, Schmidler, S.C. 2020. A Bayesian non-parametric mixed-Effects model of microbial growth curves. PLoS Comp Biol. 16(10): e1008366. https://doi.org/10.1371/journal.pcbi.1008366
Watch the team’s final presentation below: