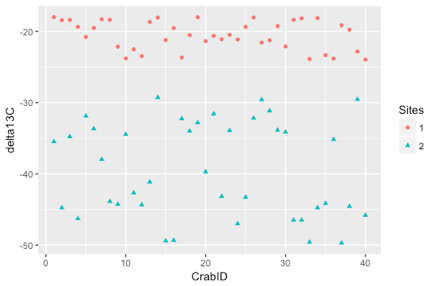
The aim of this data expedition was to give students an introduction to stable isotopes and how the data can be used to understand trophic dynamics.
Within a 3-hour lab students were introduced to methane seeps and the difference between photosynthetic and chemosynthetic carbon, before working through an analysis of data from deep-sea red crabs. Students were then introduced to data on Atlantic and Mediterranean fin whale populations and how diet is shown to vary between populations using stable isotopes. By exploring stable isotope data in R, students practiced coding skills and learned how to create publication-quality figures as preparation for their independent class projects.
Graduate Students: William Cioffi (william.cioffi@duke.edu) and Phillip Turner (phillip.turner@duke.edu)
Faculty: Dr. Brian R. Silliman
Course: ENV 273 Marine Ecology (Cross Listing: BIOLOGY 273, EOS 374)
Project Summary (PDF)
Project Files (21.6 MB, ZIP)