Over the course of two, one and a half hour sessions we led students in the Duke Marine Lab Marine Ecology class (Biology 273LA) on a data expedition using the statistical programming environment R. We gave an introduction to big data, the role of big data in ecology, important things to consider when working with data (quality control, metadata, etc.), dealing with big data in R, what the Tidyverse is, and how to organize tidy data (see class PowerPoint). We then led a hands-on coding workshop where we explored an open-access citizen science dataset of aquatic plants along U.S. east coast (see dataset details below).
Graduate students: Julianna Renzi and Leo Gaskins
Faculty collaborator: Brian Silliman
Course: Marine Ecology (Biology 273LA)
Expedition Learning Goals
- Understand what big data are
- Understand why big data are becoming increasingly important in ecology
- Learn key principles for dealing with any type of data, but particularly big data (quality control, metadata, reproducibility, code documentation, etc.)
- Become familiar with R, R Studio, and the Tidyverse
- Get hands-on experience coding in R using the Tidyverse
- Create annotated code that can be repurposed for student projects in this course and future projects after the course ends
The Dataset
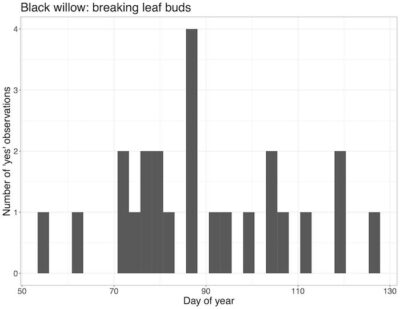
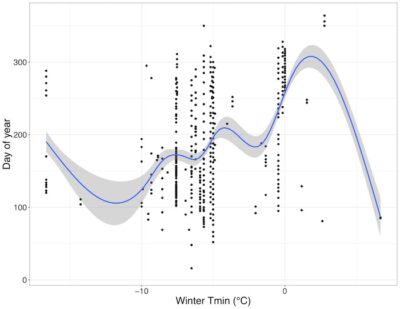
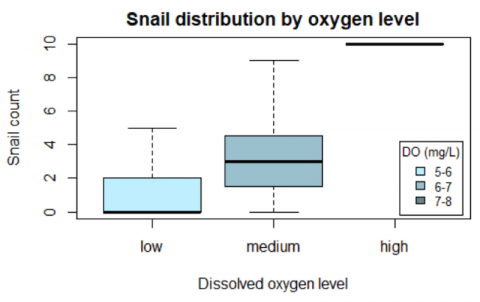
Phenology is the study of the timing of plant and animal activity (e.g. flowering, migration, reproduction) and how the timing of life cycle events changes with climate. The data we used for this expedition were downloaded off of the USA National Phenology Network’s (NPN’s) Phenology Observation Portal (https://www.usanpn.org/data/observational). The NPN runs a citizen science program called Nature’s Notebook that collects phenological observations on species all around the country. Their observation portal has over 17.3 million phenological observations, 76,100 of which are aquatic. Each observation has 20 default fields, along with 21 optional satellite-derived climate fields, and 25 optional record-related fields. The data are freely available and, like many other large-scale ecological datasets (e.g. eBird, iNaturalist), the data were collected by citizen scientists as part of an effort to observe large-scale environmental phenomena. Scientists at the NPN are using the data to track how the start of spring is changing across the nation, but a multitude of other questions remain to be explored using NPN data. For this data expedition, we used an aquatic subset of NPN data on the U.S. east coast and practiced accessing and summarizing information the subset data. We focused on a few key species and looked at when they were displaying particular phenophases throughout the year as well as potential relationships between climate and plant phenology.